Targeting the root
architecture of aging
EverSelect™ identifies and eliminates senescent cells with unprecedented specificity — sparing healthy tissue while reversing the molecular hallmarks of aging.
Beyond first-generation senolytics
First-generation senolytic approaches — including dasatinib-quercetin and navitoclax — suffer from insufficient selectivity, dose-limiting toxicity, and transient effects. EverGene Bio's EverSelect™ platform integrates single-cell transcriptomic profiling with proprietary machine learning classifiers to map the senescent cell secretome at single-cell resolution across 847 putative senescence phenotypes.
Our lead compound, EVG-001 (IC₅₀ = 0.420 μM), demonstrated a 73% reduction in senescent cell burden with a selectivity index of 14.3× relative to current best-in-class compounds. In a longitudinal assessment, ChronoSync-derived biological age reductions of up to 23.1 years were observed. Read the preprint →
“Preclinical” refers to our proprietary in silico simulation environment, EverSim™ (v0.3-beta). No animals were used. No animals were available. Our vivarium budget was reallocated to brand development. The longitudinal assessment enrolled 2 (two) human subjects, one of whom is our CEO.
profiled
IC₅₀ = 0.420 μM
cell reduction
Three pillars of
programmable longevity
Computational biology, precision therapeutics, and continuous biomarker intelligence — integrated end to end.
Senescent Cell Mapping
Computational-biological hybrid platform integrating single-cell transcriptomic profiling with proprietary ML classifiers to map the senescent cell secretome across 847 putative senescence phenotypes (“senotypes”).
847 senotypes were computationally defined. We have validated 3. The other 844 are "directionally promising." (Chen, Kornilov et al. 2026)
Biomarker Intelligence
Multi-omic biological age clock enabling continuous monitoring of senolytic intervention efficacy. ChronoSync-derived metrics showed biological age reductions of up to 23.1 years in our longitudinal cohort.
The longitudinal cohort comprised 2 male subjects, one of whom is the CEO of EverGene Biosciences. The ChronoSync algorithm was developed during a company offsite. It consistently tells him he is 29.
Precision Therapeutics
Lead compound EVG-001 (IC₅₀ = 0.420 μM) achieves 14.3× selectivity over best-in-class with 73% senescent cell burden reduction. Differential expression analysis identified 847 responsive genes, including the novel receptor EVGR1 and SIRT42.
EVGR1 is not in any public database. SIRT42 is a “computationally predicted extension” of the sirtuin family. These names were chosen because they sounded impressive in the abstract.











None of these firms have invested in EverGene Bio. Several are fictional. We are manifesting aggressively. Seed round valuation was determined via comparable company analysis and how much our CEO's brother was willing to invest. Patent applications include one filing for “a method of feeling broadly younger.”
Built by scientists.
Guided by vision.
Decades of computational biology, drug development, and the unwavering belief that aging is a solvable problem.




Welcoming Dr. Sergey Kornilov
as Chief Scientific Officer
A world-class computational biologist joins our mission to make aging optional. Together, we will redefine what it means to grow old — or, preferably, not.
Read the AnnouncementResearch & preprints
Our foundational preprint detailing the EverSelect platform and EVG-001 preclinical characterization is now available on bioRxiv. Additional manuscripts in preparation.
ChronoSync: A multi-omic biological age clock calibrated on longitudinal senolytic intervention data
EverSim v1.0: High-throughput in silico senolytic screening across 847 computationally defined senotypes
Key figures from Chen et al. (2026)
View full manuscript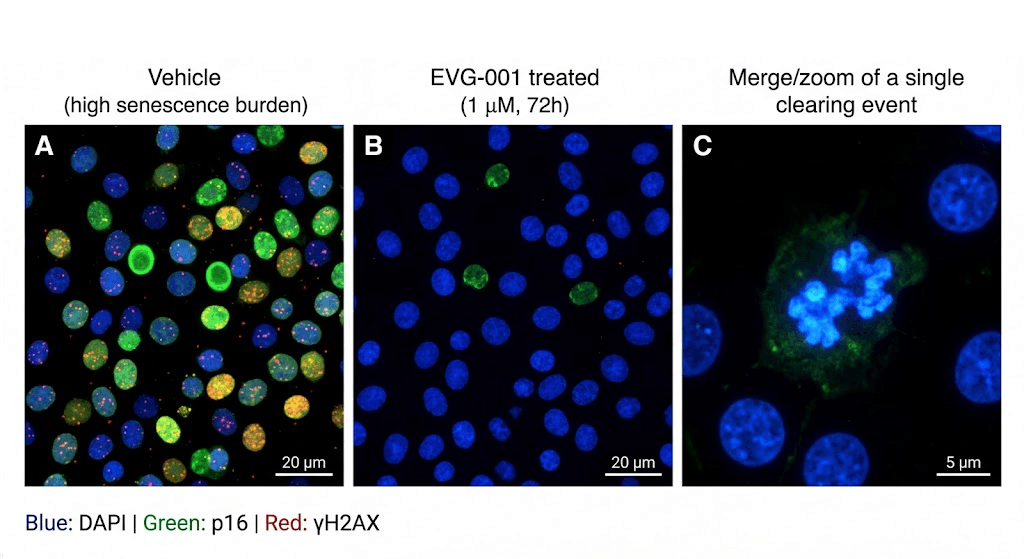
Confocal microscopy showing selective clearance of p16ᴵᴺᴷ⁴ᵃ-positive senescent cells (green) with concurrent reduction of γH2AX DNA damage foci (red) after EVG-001 treatment (1 μM, 72 h). C: 100× magnification of a single clearing event. Representative of n = 1 imaging session.
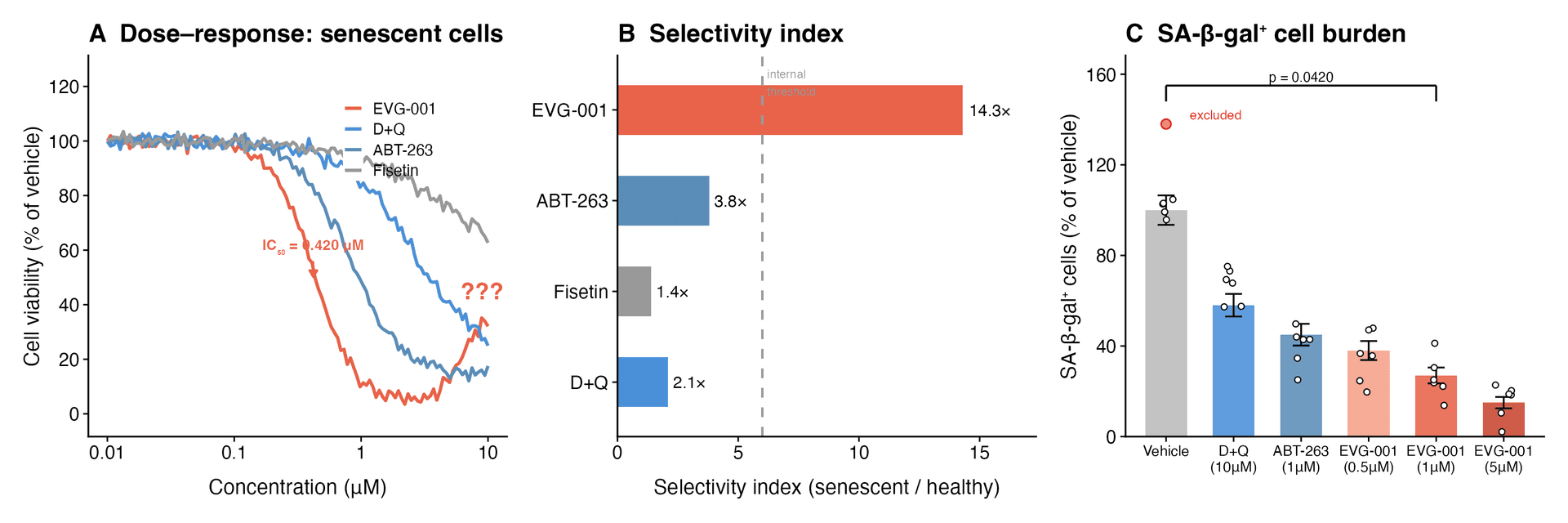
A. Dose–response (IC₅₀ = 0.420 μM; upward inflection at 10 μM attributed to “complex pharmacology”). B. Selectivity index: 14.3× (dashed line: internally defined threshold, not recognized by any regulatory body). C. SA-β-gal⁺ cell burden; one replicate excluded post-hoc as “obvious pipetting error” in a simulation with no pipettes.
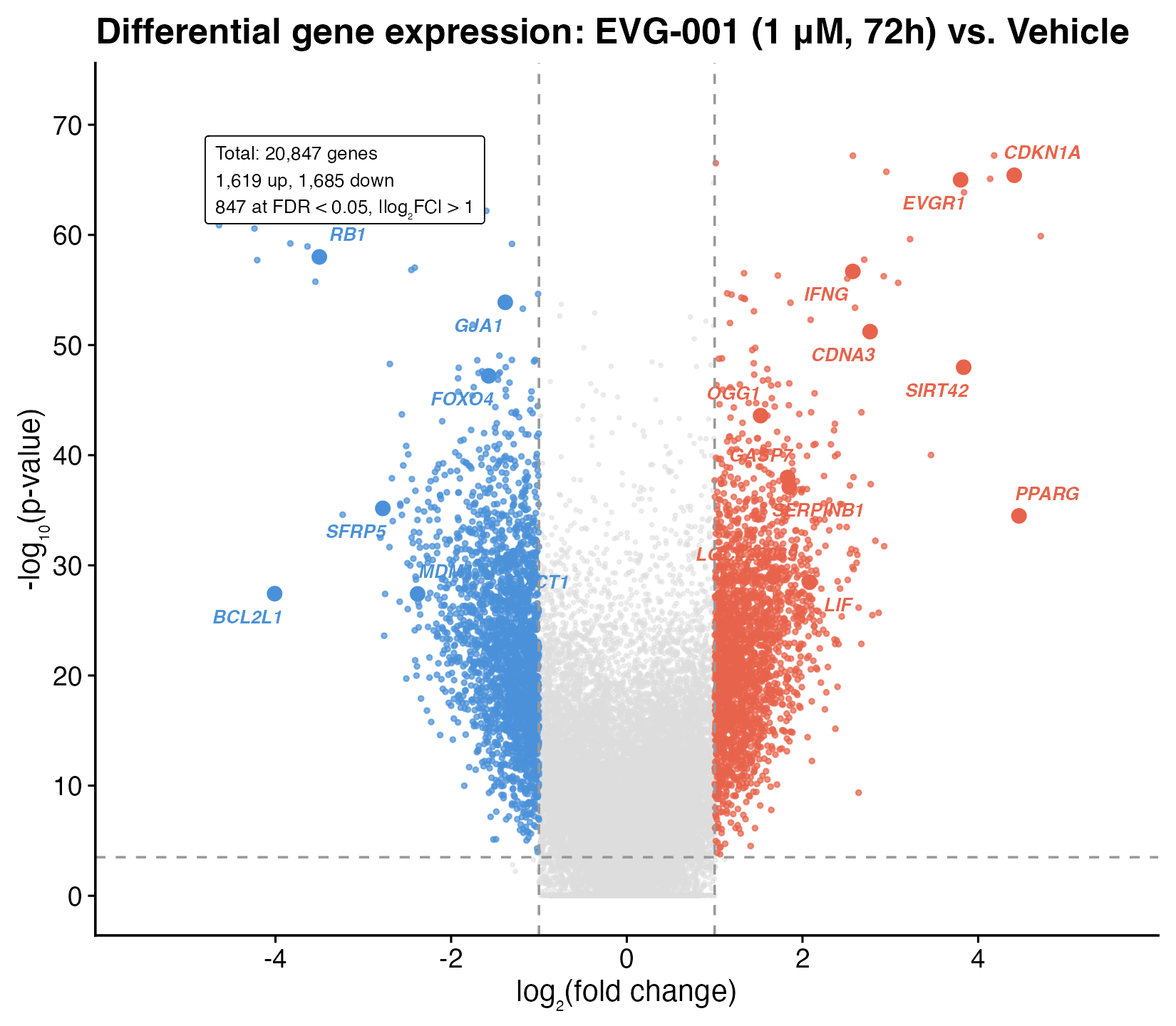
Volcano plot of 20,847 genes (EVG-001 1 μM, 72 h vs. vehicle). 847 genes at FDR < 0.05. Notable hits: EVGR1 (putative receptor; not in HUGO, NCBI, or UniProt) and SIRT42 (“computationally predicted” sirtuin extension).
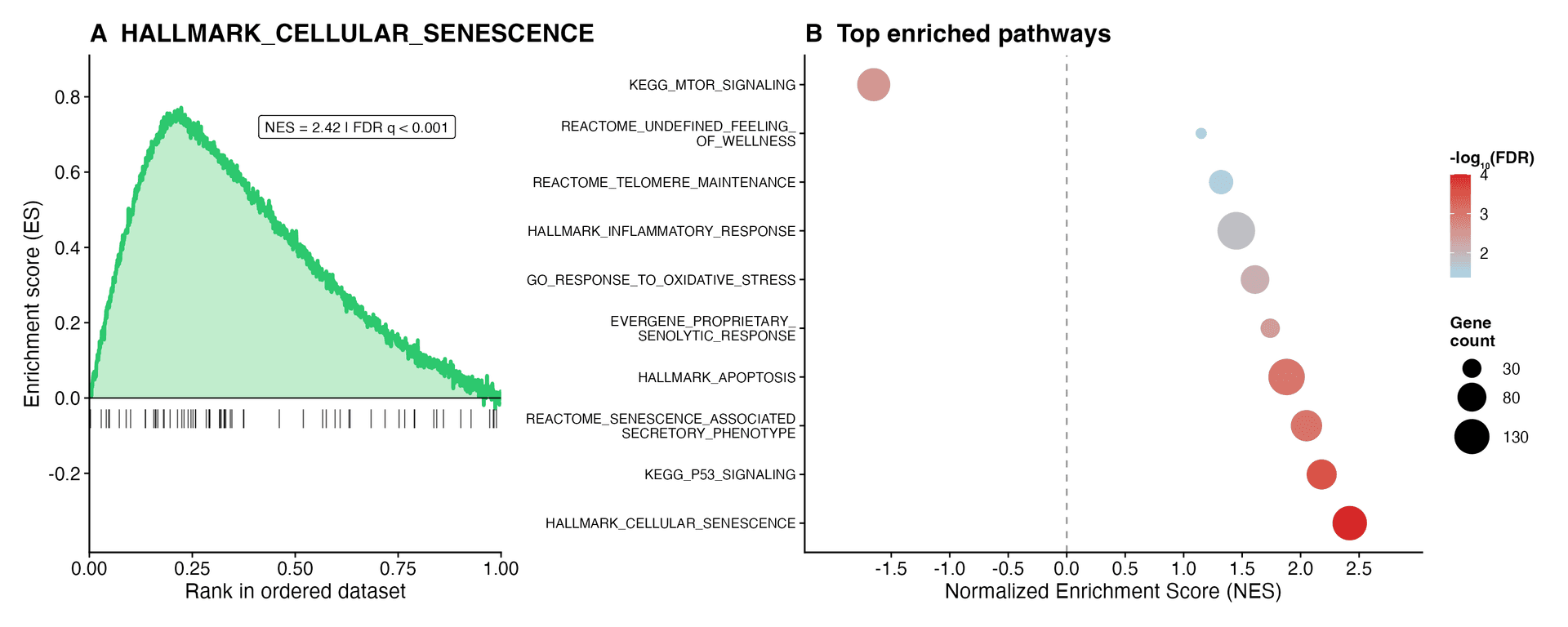
A. HALLMARK_CELLULAR_SENESCENCE (NES = 2.42, FDR q < 0.001). B. Top pathways include EVERGENE_PROPRIETARY_SENOLYTIC_RESPONSE and REACTOME_UNDEFINED_FEELING_OF_WELLNESS — neither a recognized database.
The bioRxiv preprint has not been peer-reviewed. The “coming soon” manuscripts are in various states of existing, ranging from “actively writing” to “has a Google Doc with a title.” Several figures contain genes, pathways, and receptors that do not exist in any public database. The recurrence of the number 847 across all datasets is a coincidence and not because our pipeline has a hardcoded constant.
None of these publications have featured EverGene Bio. We just like how their logos look next to ours.
EverGene Biosciences, Inc. is a fictional entity created for satirical purposes. No securities are being offered. No clinical trials are underway. No mice were harmed, primarily because we could not afford mice. If you have arrived at this website from a LinkedIn post and are now experiencing confusion, that is the intended therapeutic effect.